-
Notifications
You must be signed in to change notification settings - Fork 3
Clone sizes issue #18
Description
Hi @rschenck
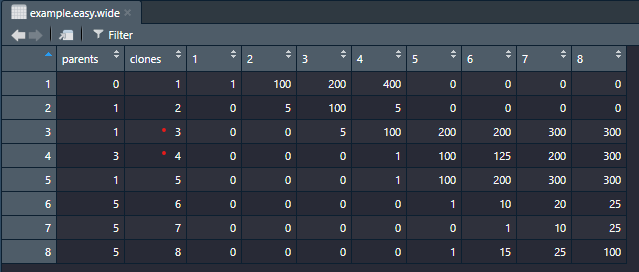
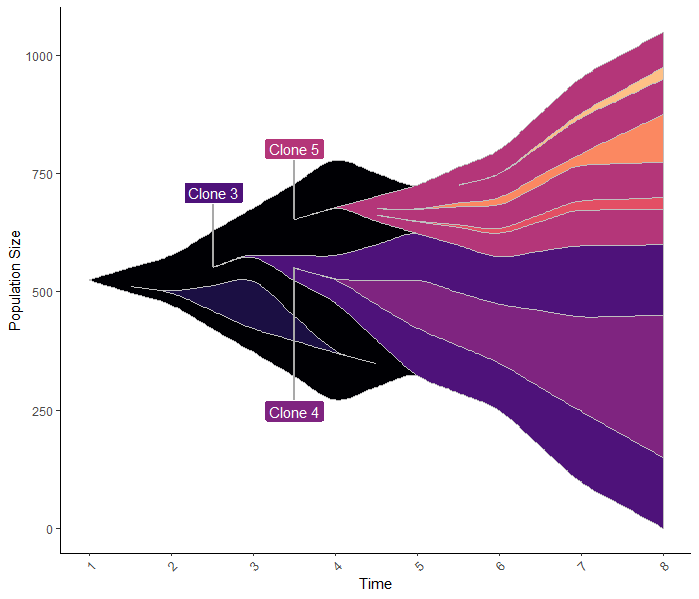
Nice library. A bit confused about how your package deals with clone sizes though. Looks like it blindly stacks up all clones as if they are independent (almost never the case). Below is the plot and df generated from your internal data. Clone 4 is a child of clone 3 and in the dataframe they both have the size of 300 but the plot obviously blows up the parental clone to 600. Usually clone size is derived from VAF/CF and cellularity data so having that data and clonal structure one could immediately put the evoseq df together without adjusting for 'stacking issue' in your library. So the whole tumor size at time point 8 should be just the sum of two dominating clones (clone 3 and clone 5) which is 600 according to df, but it's over 1000 in the plot. I doubt it's a feature, or is it? Thoughts?
Thanks!