2nd CAMI Toy Human Microbiome Project Dataset has been chosen as a testing dataset. It consists of 49 samples of simulated metagenome data from five different body sites of the human host (10 gastrointestinal tract, 10 oral cavity, 10 airways, 10 skin, 9 urogenital tract). Follow the instructions:
Step 0:
Download dependencies:
- CamiClient (
curl -O https://data.cami-challenge.org/camiClient.jar). - Python3 and packages numpy, pandas, scipy and seaborn.
Step 1:
Execute download script. You will get five folders (Airways, Skin, Oral, Gastro, Uro) with metagenome files, each inside it`s own folder /short_read/2017.12.04_18.56.22_sample_<id>/reads. All files have the same name anonymous_reads.fq.gz, so in order to receive more visual image, run this script for renaming files.
Step 2:
Run MetaFast by executing following command (full script):
java -jar ./metafast.jar -t matrix-builder -k 23 -i <List_of_CAMI_files> -b 5 -l 1200
Several attempts have been made to select the best parameters values: k-mer size is 23, minimum sequence length to be added to a component is 1200 and maximal frequency for a k-mer to be assumed erroneous is set to 5.
As a result, the Bray-Curtis dissimilarity matrix and corresponding image file with heatmap and dendrogram are produced.
The image shows five distinct clusters (saved in workDir/matrices/dist_matrix_<date>_<time>_heatmap.png) :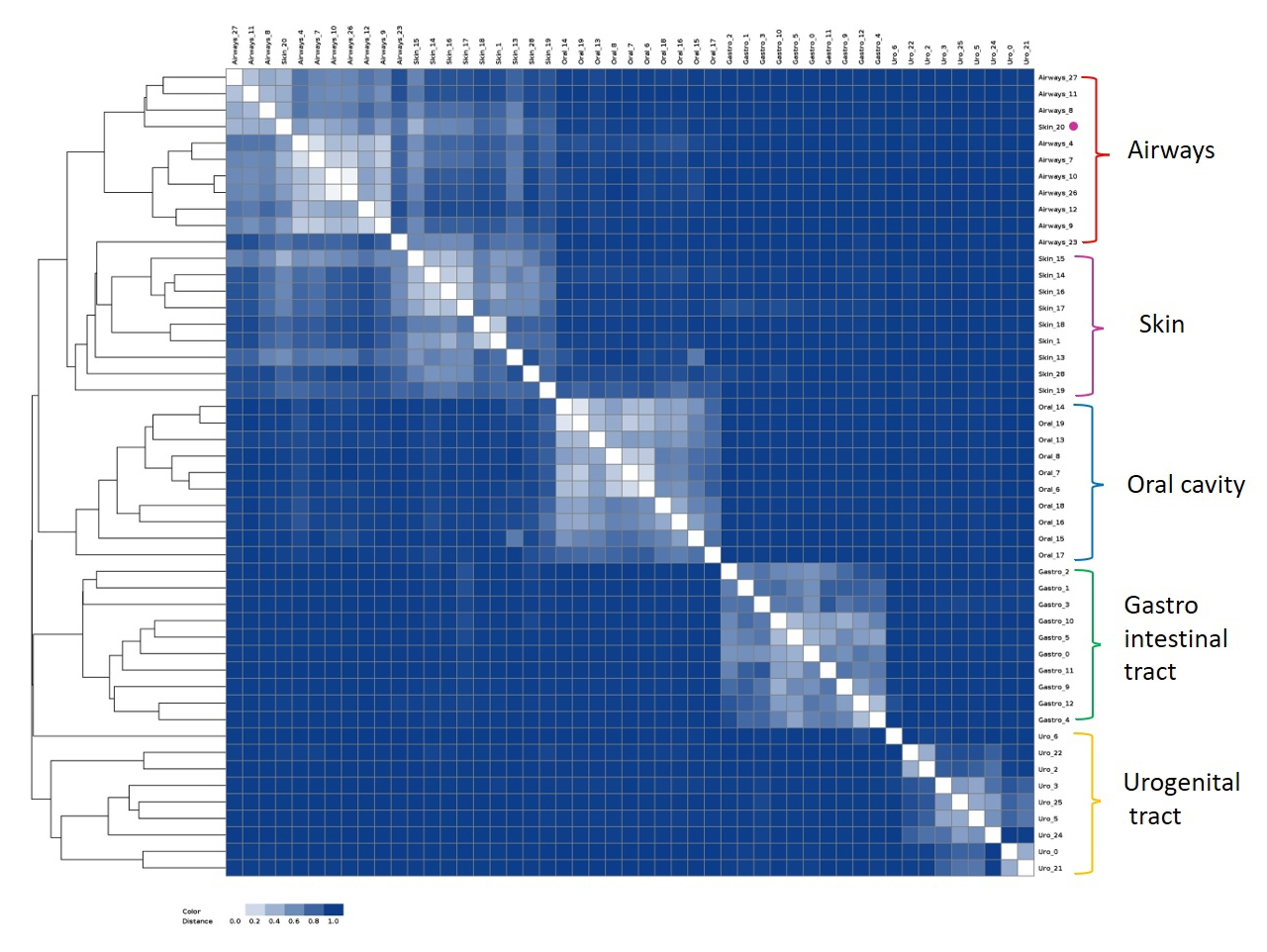
Taxonomic clustering
For comparison, files with taxonomic data taxonomic_profile_<id>.txt downloaded for each group by running this script. Then dissimilarity matrix using Bray-Curtis metric was evaluated, and with seaborn.clustermap an analogous image file with heatmap and dendrogram was produced:
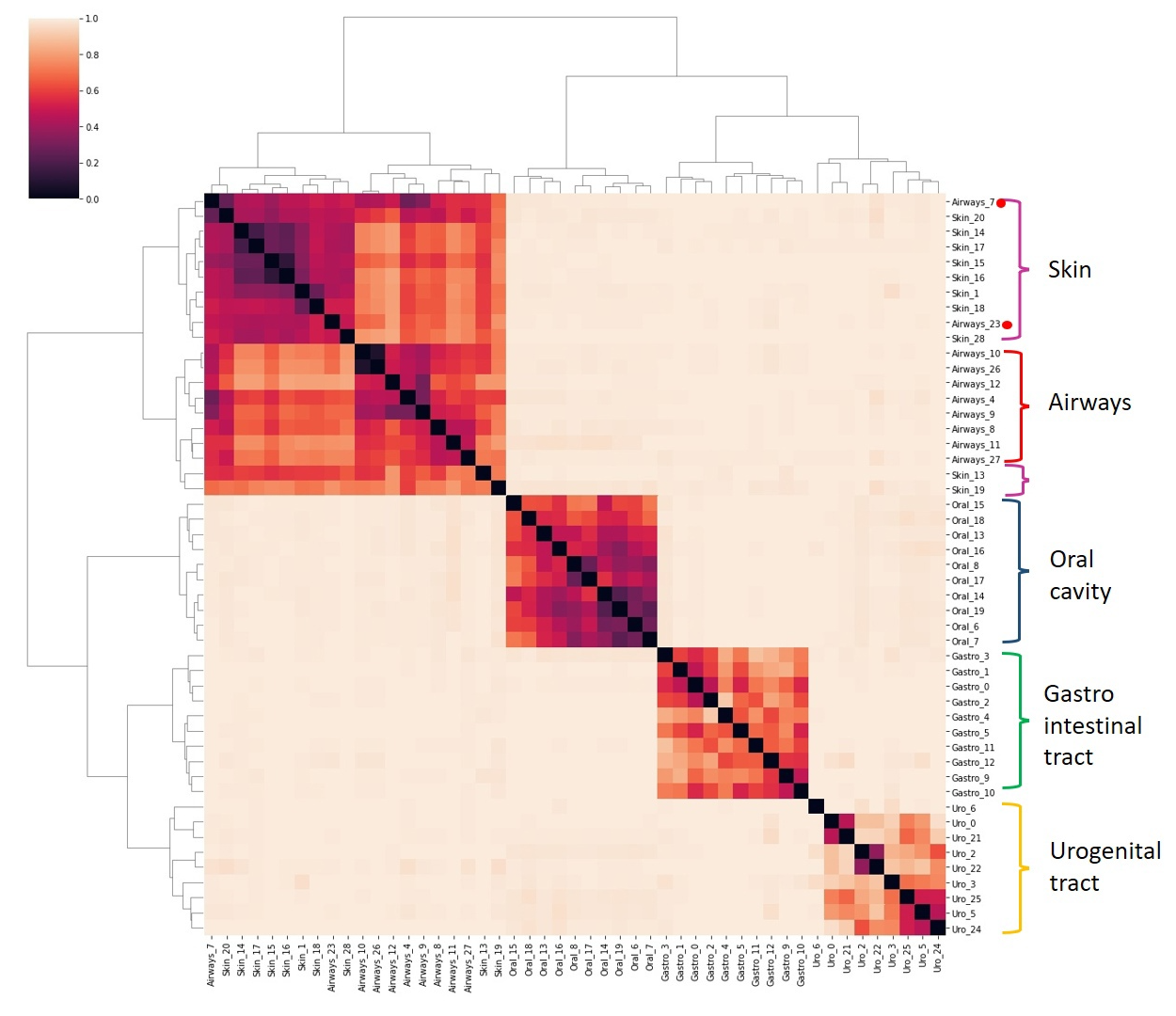 Here you can see a Python3 script.
Here you can see a Python3 script.
As we can see, the results of the two methods are quite similar. Both images show a clear distinction between oral cavity, gastrointestinal tract, and urogenital tract groups. In contrast, both approaches faced difficulties distinguishing between airways and skin groups, but MetaFast demonstrated a more evident differentiation.