snputils is a Python package designed to ease the processing and analysis of genomic datasets, while handling all the complexities of different genome formats and operations very efficiently. The library provides robust tools for handling sequencing and ancestry data, with a focus on performance, ease of use, and advanced visualization capabilities.
Developed in collaboration between Stanford University's Department of Biomedical Data Science, UC Santa Cruz Genomics Institute, and more collaborators worldwide.
Note: snputils is under active development. While the core API is stabilizing, we are continuously adding features, optimizing performance, and expanding format support.
Basic installation using pip:
pip install snputilsOptionally, for GPU-accelerated functionalities, install the package with the [gpu] extra:
pip install 'snputils[gpu]'snputils is designed to be user-friendly and intuitive, with a simple API that allows you to quickly load, process, and visualize genomic data. For example, reading a whole genome VCF file is as simple as:
import snputils as su
snpobj = su.read_snp("path/to/file.vcf.gz")Similarly, reading BED or PGEN filesets is straightforward:
snpobj = su.read_snp("path/to/file.pgen")Working with ancestry files, performing processing operations, and creating visualizations is just as straightforward. See the demos directory for examples.
snputils aims to provide the fastest available readers and writers for various genomic data formats:
- VCF: Support for
.vcfand.vcf.gzfiles - PLINK1: Support for
.bed,.bim,.famfilesets - PLINK2: Support for
.pgen,.pvar,.psamfilesets - Local Ancestry: Handle
.msplocal ancestry format - Admixture: Read and write
.Qand.Pfiles
-
Basic Data Manipulation
- Filter variants and samples, correct SNP flips, and filter out ambiguous SNPs
- Compute cohort allele frequency and ancestry-specific allele frequencies via
SNPObject.allele_freq(...)or in streaming for memory efficiency withsnputils.stats.allele_freq_stream(...) - Standardized querying across genotype, local ancestry, global ancestry, and IBD data
-
Dimensionality Reduction
- Standard PCA with optional GPU acceleration
- Missing-data PCA (mdPCA)
- Multi-array ancestry-specific MDS (maasMDS)
-
Population Genetic Statistics
- Compute
$D$ ,$f_2$ ,$f_3$ ,$f_4$ , the$f_4$ -ratio, and$F_{ST}$ (Hudson and Weir-Cockerham) - Includes block jackknife standard errors and optional ancestry masking
- Compute
-
Identity-by-Descent (IBD) & Relatedness
- Read
hap-IBDandancIBDoutputs into a unified format - Fast filtering and ancestry-restricted segment trimming using local ancestry
- Read
-
Admixture Analysis & Simulation
- Admixture Mapping: Locus-by-locus regression of local ancestry dosage on traits
- Simulation: Lightweight haplotype-based simulation of admixed mosaics from real founder haplotypes
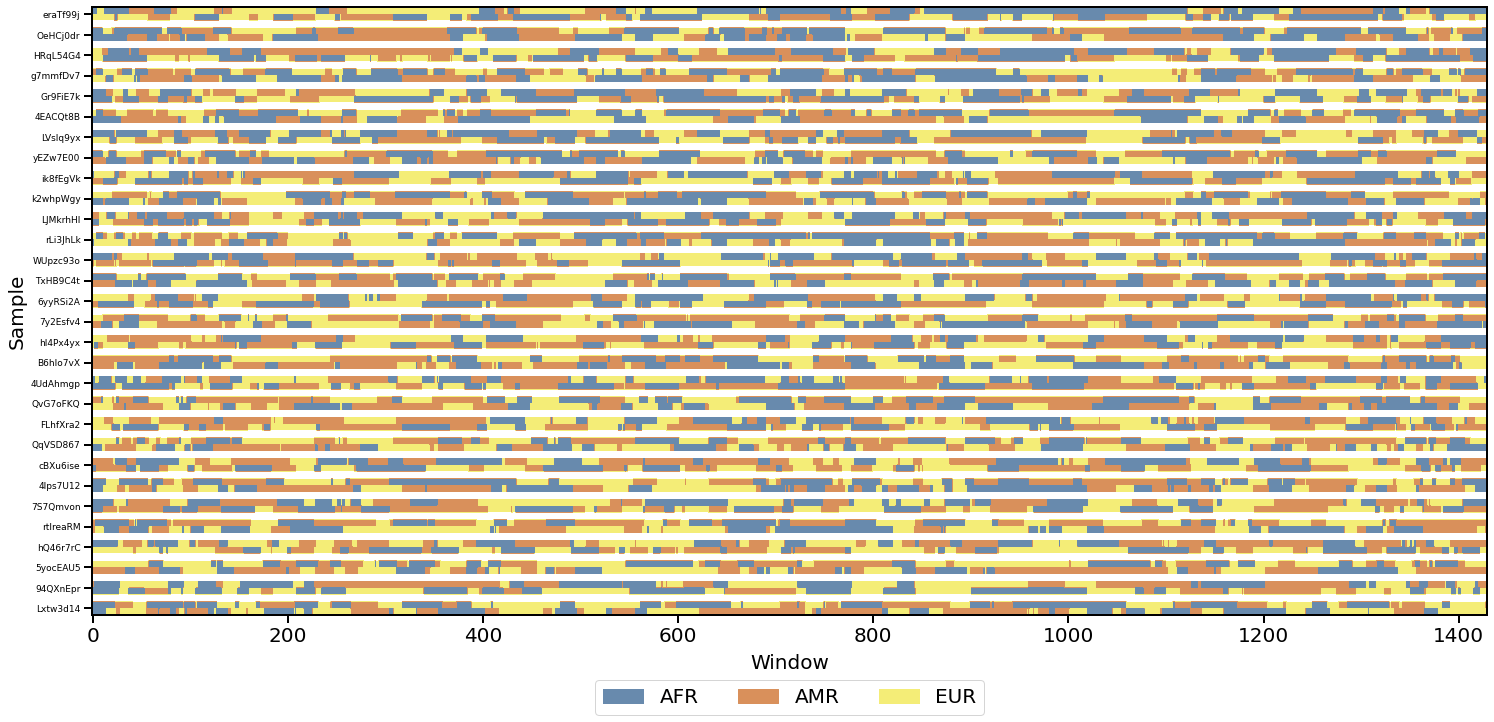
- Interactive global ancestry bar plots
- Detailed scatter plots of PCA, mdPCA, and maasMDS
- Admixture mapping Manhattan plots
- Local ancestry visualization
- Chromosome painting (with Tagore)
- Dataset-level
- Fast file I/O through built-in methods or optimized wrappers (e.g., Pgenlib for PLINK files)
- Memory-efficient operations using NumPy and Polars
- Optional GPU acceleration via PyTorch for computationally intensive tasks
- Support for large-scale genomic datasets through efficient memory management
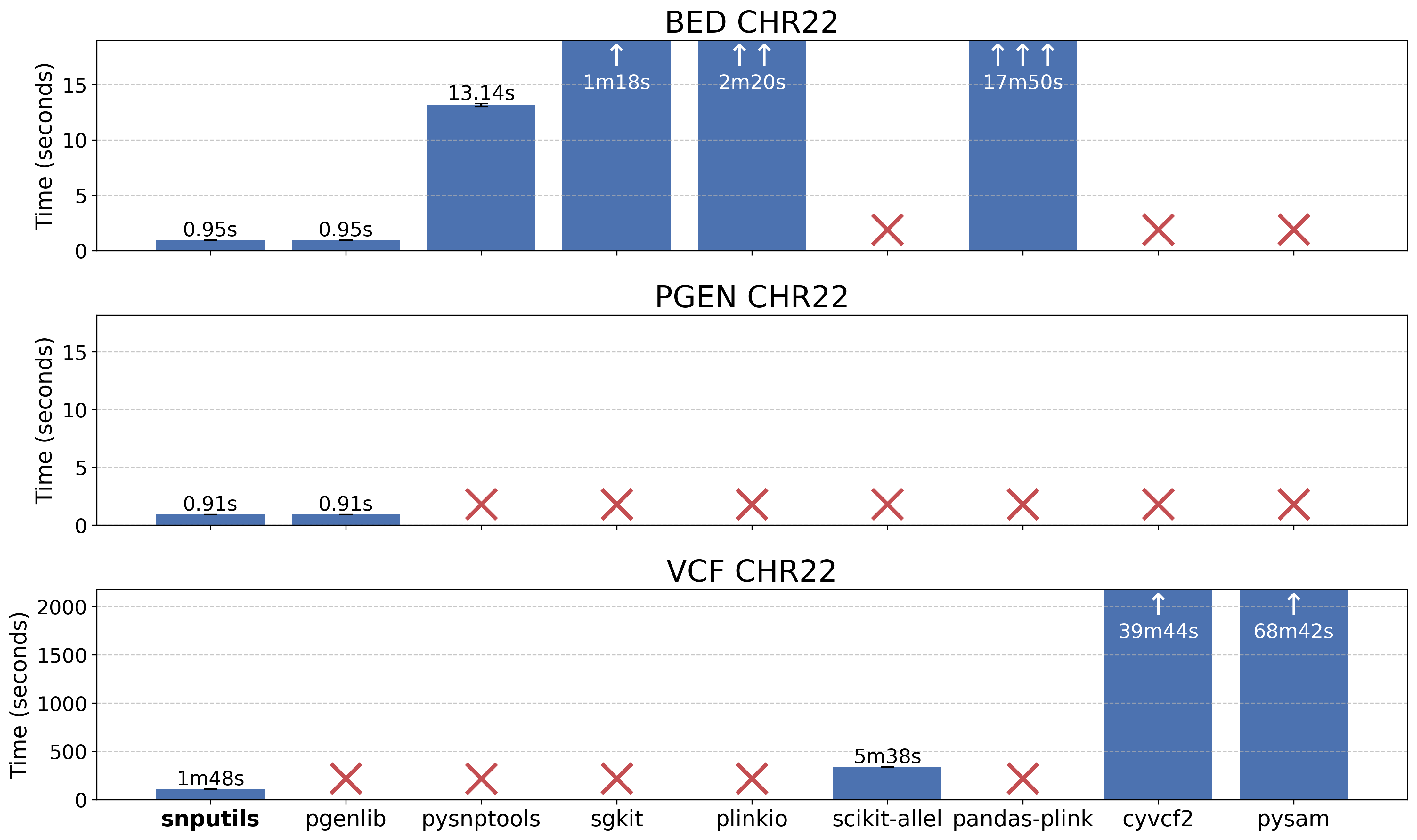
Our benchmark demonstrates superior performance compared to existing tools:
Reading performance comparison for chromosome 22 data across different tools. See the benchmark directory for detailed methodology and results.
The snputils package is continuously updated with new features and improvements.
- Documentation: Comprehensive API reference at docs.snputils.org.
- Examples & Tutorials: Check out our interactive notebooks in the demos directory.
- Issues & Community: Report bugs, ask questions, or request features via GitHub Issues.
If you use snputils in your research, please cite our paper:
@article{snputils2026,
author = {Bonet, David and Comajoan Cara, Marçal and Barrabés, Míriam and Smeriglio, Riccardo and Agrawal, Devang and Aounallah, Khaled and Geleta, Margarita and Dominguez Mantes, Albert and Thomassin, Christophe and Shanks, Cole and Huang, Edward C. and Franquesa Monés, Marc and Luis, Aina and Saurina, Joan and Perera, Maria and López, Cayetana and Sabat, Benet Oriol and Abante, Jordi and Moreno-Grau, Sonia and Mas Montserrat, Daniel and Ioannidis, Alexander G.},
title = {{snputils}: A High-Performance {Python} Library for Genetic Variation and Population Structure},
year = {2026},
month = feb,
doi = {10.64898/2026.02.28.708618},
url = {https://www.biorxiv.org/content/10.64898/2026.02.28.708618},
journal = {bioRxiv},
publisher = {Cold Spring Harbor Laboratory},
}We would like to thank the open-source packages that make snputils possible.