Neural networks have emerged as powerful tools to understand the functional relationship between genomic sequences and various biological processes. However, current practices of training and evaluating models on genomic sequences may fail to account for the widespread homology that permeates the genome. Homology spanning train-test data splits can result in data leakage, potentially leading to overestimation of model performance and a reduction in model reliability and generalizability.
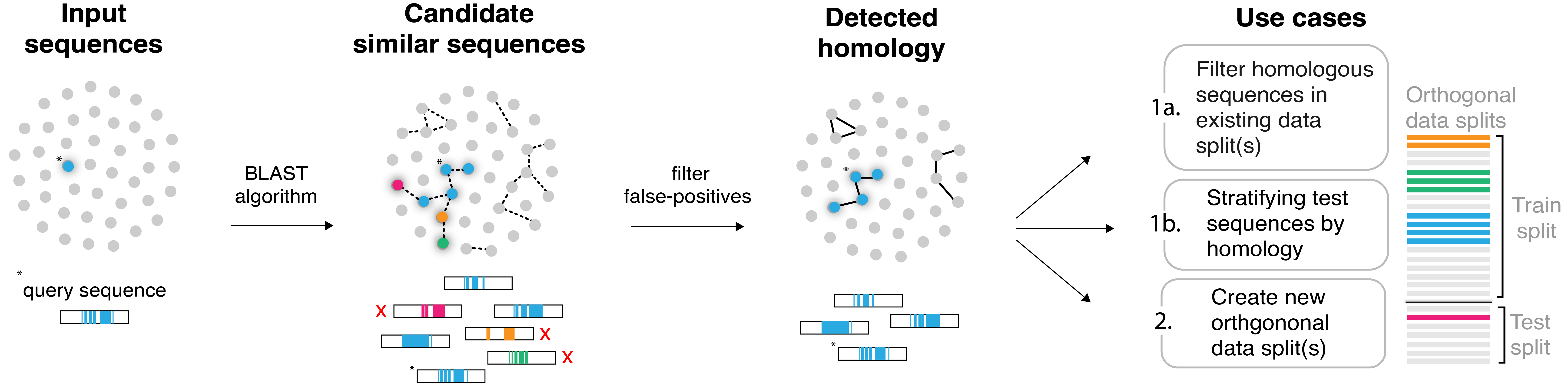
hashFrag is a scalable command-line tool to help users address homology-based data leakage during model development. The general workflow involves identifying “candidate” pairs of sequences exhibiting high similarity with BLAST, filtering these candidates based on a specified similarity threshold, and then using the resulting homology information to mitigate the potential occurrences of data leakage in existing or newly-created splits.
Full documentation is available on Read the Docs.
Check out our preprint on bioRxiv titled, "Detecting and avoiding homology-based data leakage in genome-trained sequence models", for more details.